Contact : Franz Bruckert, Marianne Weidenhaupt
Adsorption of protein on material surfaces is of widespread importance in the fields of cellular biology, pharmacology and medicine. Although protein adsorption and aggregation upon contact with material surfaces has been studied for a long time, the role of the material surface itself has not been examined in detail.
We use sensitive biochemical (BCA) and biophysical (SPRi, ATR-FTIR) and microscropy techniques (fluorescence imaging AFM, TEM,) to study protein adsorption and aggregation kinetics in a quantitative and qualitative way.
Our analysis focuses both on the protein and on the material and we explore the effects of different physico-chemical parameters of the material and its environment on protein aggregation.
The mathematical modeling of protein adsorption and aggregation on material surfaces is another aspect of our research.
We have established a 96-well based protocol using insulin as a model protein and have shown that insulin adsorbs on hydrophobic surfaces and that this adsorption is accompanied by a conformational change involving alpha-helix to beta-sheet transitions. Once a sufficient amount of protein is accumulated on the surface, aggregation nuclei are formed, allowing massive aggregation of insulin into amyloid fibers.
We have demonstrated that the aggregation nuclei are created solely on the surface of the material and not in solution, making aggregation strictly dependent on the hydrophobic surface.
In order to improve our understanding of the molecular mechanisms at the material surface, we use amyloidogenic peptides of insulin and bacterial chaperones, known to either accelerate or inhibit its amyloid aggregation on materials (Figure 1).
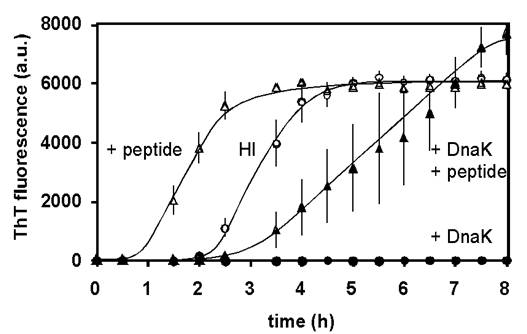
Figure 1. Effect of amyloidogenic peptide and chaperones on aggregation kinetics
Publications:
Ballet et al. 2012, accepted in Biochemistry http://dx.doi.org/10.1021/bi201457u
Collaborations:
Horiba Scientific
Bekton Dickinson, Pharmaceutical Systems, France
Laboratoire Français du Fractionnement et des Biotechnologies, LFB
CEA Grenoble /iRSTV/ Chimie et Biologie des Métaux/Équipe Amyloid Fibres: From Foldopathies to NanoDesign)



